In the postgenomic era, the numerous and diverse noncoding RNA species once dismissed as “junk RNA” are increasingly seen as treasure. Noncoding RNAs, we now know, have diverse functions in health and disease.
Some in the field believe that we have only started to appreciate the riches of noncoding RNA. The ultimate jewels? They may prove to be previously hidden connections with cancer.
....
Prostate Cancer and Noncoding RNA
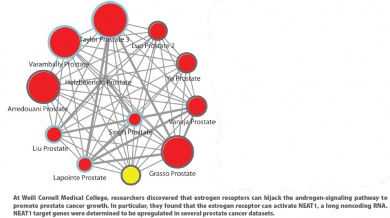
Given the roles played by ncRNAs in a host of biological processes, it is no surprise that these species also impact prostate cancer progression and therapy resistance. Nonetheless, details of the relationship between ncRNAs and prostate cancer remain to be elucidated, said Dimple Chakravarty, Ph.D., an assistant professor of pathology and laboratory medicine at Weill Cornell Medical College.
“Deregulated or aberrant expression of steroid nuclear receptors are linked with cancer progression and thus are also major targets for therapeutic intervention,” observed Dr. Chakravarty. “But specific therapies are often inadequate.
“For example, the androgen receptor [AR] plays a central role in this malignant progression. Despite the initial effectiveness of therapeutic androgen ablation, resistance inevitably develops to both first generation anti-androgen therapies and to second-generation AR-targeted therapies. The reasons for this are unclear.”
Dr. Chakravarty and colleagues wanted to better understand the role of the estrogen receptor alpha (ERα) that is expressed in prostate cancers. “Our studies identified an ERα-specific noncoding transcriptome signature. This lured us into the noncoding world,” she disclosed.
 Dr. Dimple Chakravarty, Photo credit: Weill Cornell Art & Photography
Dr. Dimple Chakravarty, Photo credit: Weill Cornell Art & Photography
Dr. Chakravarty and her collaborators, including Mark A Rubin, M.D., a professor of pathology and laboratory medicine at Weill Cornell, scrutinized a combination of chromatin immunoprecipitation (ChIP) and RNA-sequencing data. The investigators found that the most significantly overexpressed and ERα-regulated lncRNA in prostate cancer samples was a transcript called NEAT1, the nuclear enriched abundant transcript.
“Our studies utilized a battery of approaches,” detailed Dr. Chakravarty. “We used qRT-PCR and RNA-ISH to examine NEAT1 mRNA levels in prostate cancer tissue and in cell lines, and we analyzed public datasets of normal versus prostate cancer with advanced disease. Epigenetic studies demonstrated that NEAT1 is recruited to the chromatin of prostate cancer genes and contributes to an epigenetic ‘on' state.”
Dr. Chakravarty expressed excitement over these findings: “This study is the first of its kind to demonstrate transcriptional regulation of lncRNAs by an alternative steroid receptor in prostate cancer. We believe NEAT1 could serve as both a prognostic marker for aggressive prostate cancer and also a potential therapeutic target.
“Completed and ongoing studies suggest NEAT1 is a good marker for patient risk stratification and a predictor of therapy resistance. We are now exploring the possibility of knocking it out in vivo to see if there is a therapeutic benefit. It could be that targeting NEAT1 and the androgen receptor in combination may provide a unique treatment strategy for a subset of patients who have advanced prostate cancer.”
This is an excerpt of an article that appeared in Genetic Engineering & Biotechnology News. Read the full story here.
